Bay Area SCAN weekly update 3/29/23
This blog post describes data collected at 8 wastewater treatment plants in the Greater Bay Area of San Francisco, CA, including Sacramento, that are partners in the SCAN project which began in late 2020.
All samples picked up by the couriers as of 3/28/23 have been processed and their data are on the site: wbe.stanford.edu. The data from the SCAN sites are also on data.wastewaterscan.org. If you notice any bugs on the new site or have any comments about it, please email Amanda Bidwell at albidwel@stanford.edu.
COVID-19
SARS-CoV-2 and Variants
SARS-CoV-2 N gene concentrations at SCAN sites are similar to what they were two weeks ago. Concentrations are approximately between 100,000 copies per gram and 500,000 copies/g, still well above our lower detection limit of approximately 1000 copies/g. Below is a chart showing data from the San Jose plant showing similar concentrations in the N gene concentration (blue line) relative to two weeks ago. You can see some periodicity in the data over the last month.
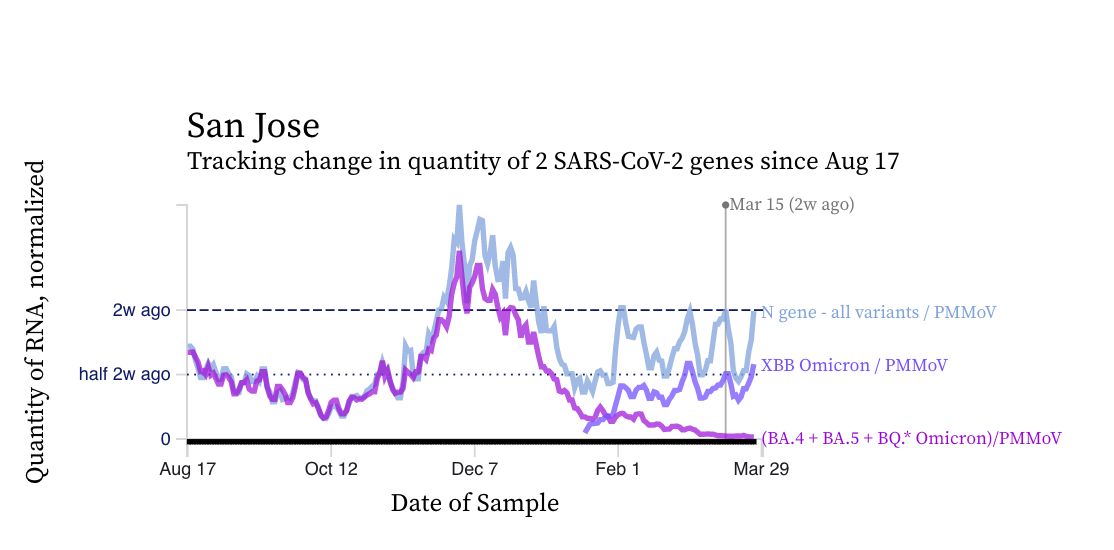
The chart shows that the mutation marker for XBB* (shown in violet) is closer to the blue line (showing the N gene, present in all variants) than the bright purple line (showing the mutation present in the BQ* and BA.5 sublineages). This indicates that most of the individuals shedding SARS-CoV-2 in the sewersheds are likely infected with and shedding XBB* rather than an other variant. We see the pattern across all the sewersheds in SCAN.
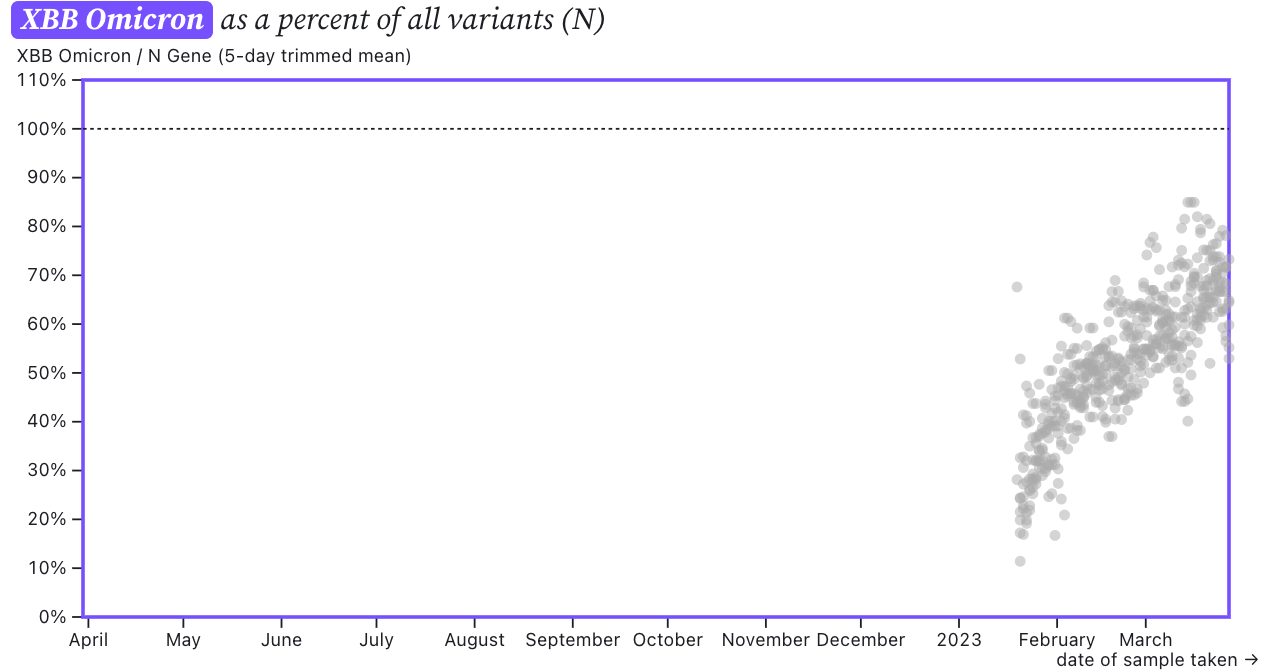
The ratio of the XBB* mutations/N across SCAN plants is shown in the chart below. As the ratio reaches 100%, it suggests that all the SARS-CoV-2 genomes in wastewater have the XBB* mutations. These data indicate that the regional average ratio XBB* mutations/N is ~75% but some places show levels as high as 80%. The trend in this percentage has been increasing. These data suggest that the majority of infections across the SCAN communities are caused by XBB or XBB.1.5. We are looking into whether this assay would detect XBB.1.16.
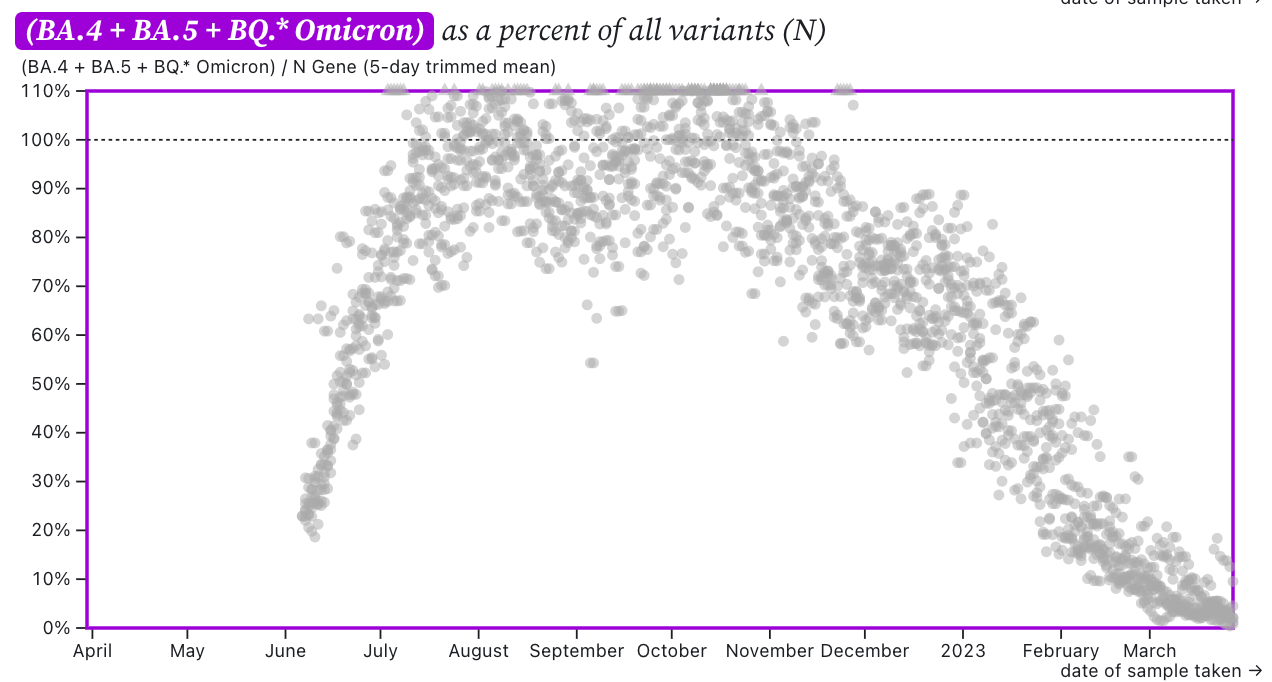
The chart below shows the ratio of the BA.5, BA.4, and BQ* mutation/N across SCAN plants. It can be read the same way as the above chart. The regional average ratio is less than 10% and has a decreasing trend.
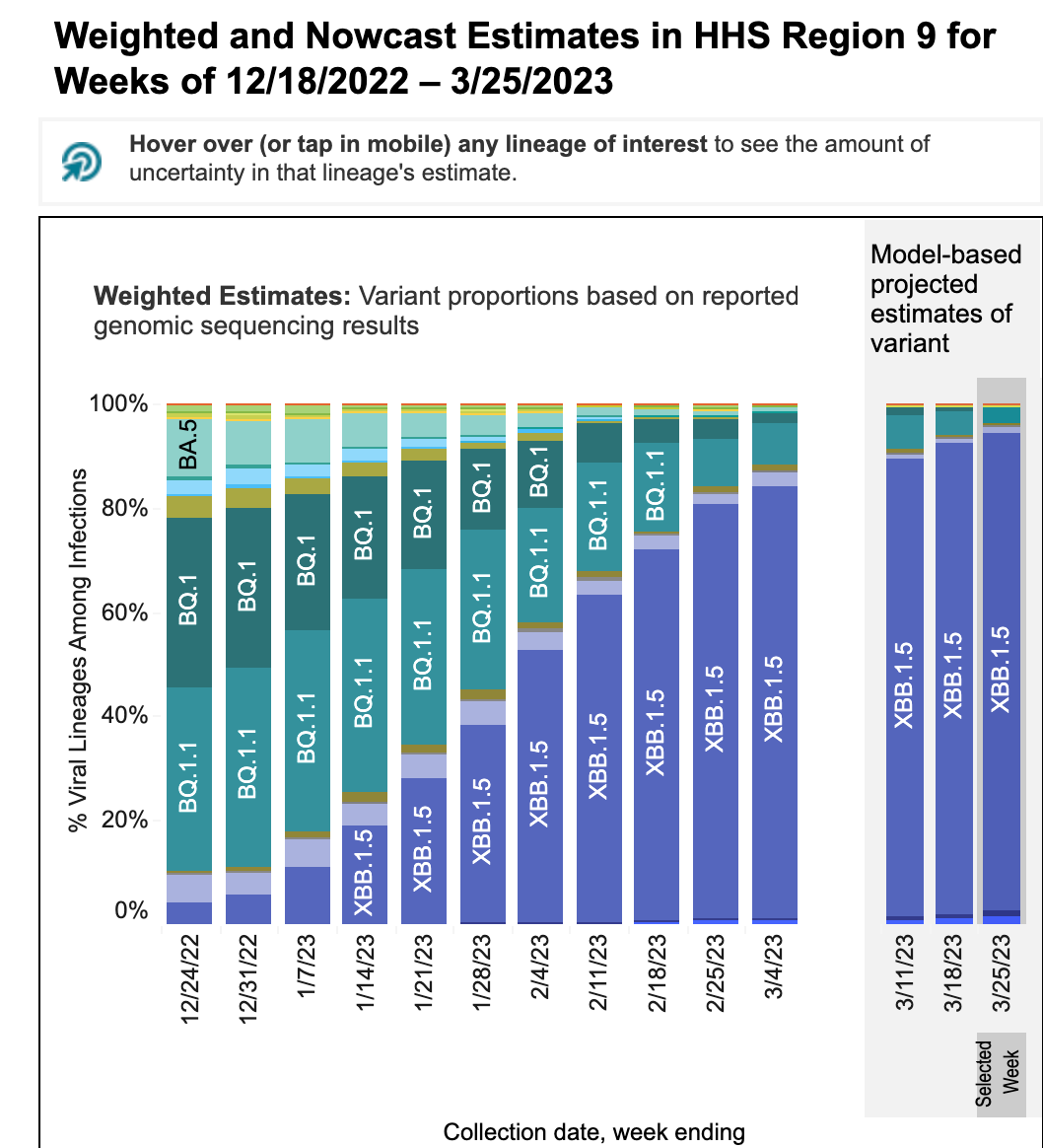
These data from wastewater solids on SARS-CoV-2 variants is consistent with the results from the CDC Nowcast for HHS region 9 (which includes CA). The nowcast suggests approximately 95% of infections in HHS region 9 are caused by XBB1.5 and <5% by BQ* (see screen shot from website below). This essentially matches the data from the SCAN plants. It is important to note that the clinical sample sequencing data used to make the CDC nowcast charts is only available as recently as a month ago (week of 3/4/23 in the chart below) and the rest of the data are model-based predictions.
Other Respiratory Viruses
IAV, IBV, RSV, and HMPV
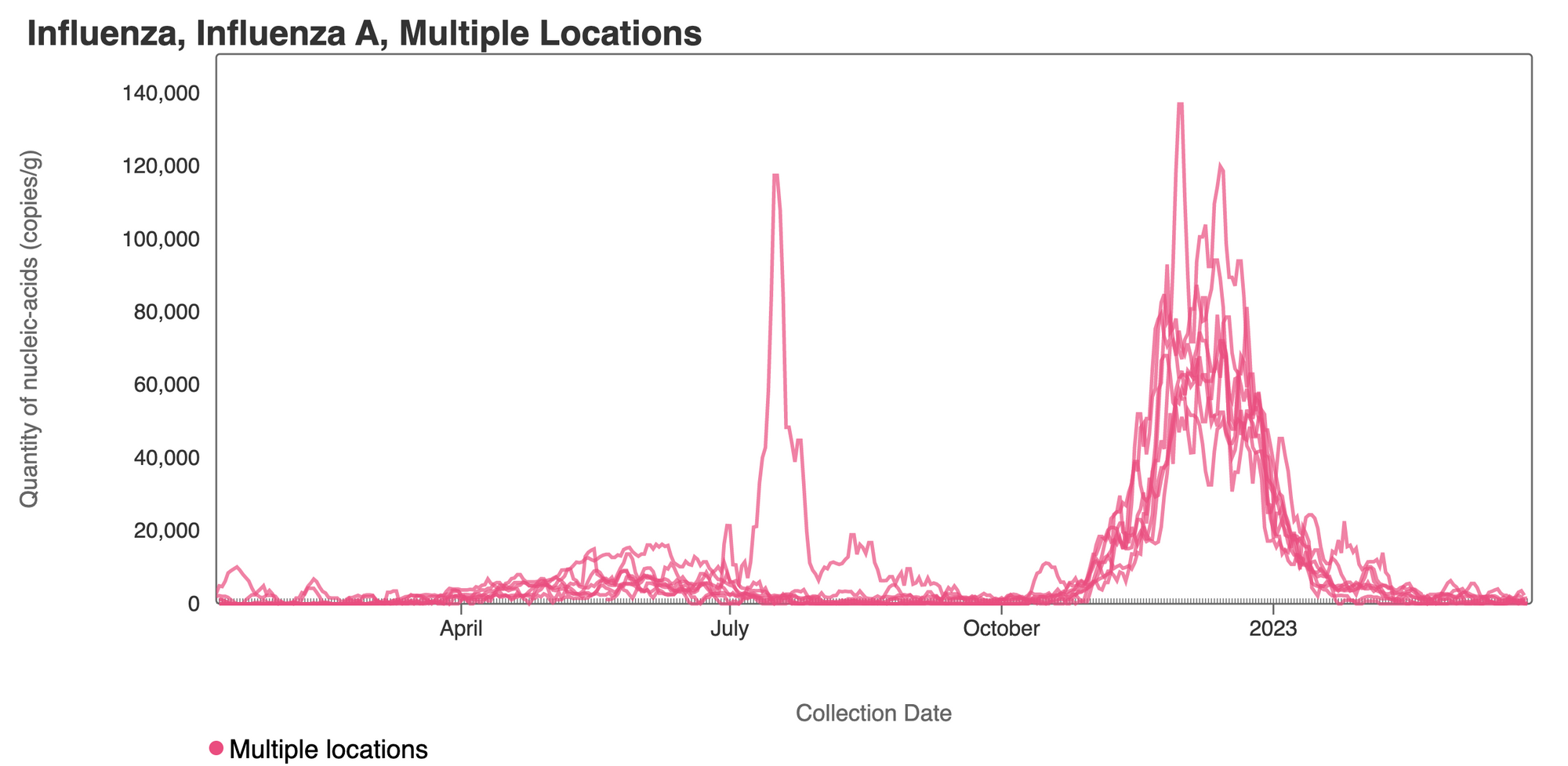
Influenza A (IAV) RNA concentrations have dropped across the plants. Concentrations are generally non-detects in samples from recent works. Influenza A RNA appears to no longer be shed into the wastewater in the SCAN plants. The link to the chart is here if you would like to interact with it.
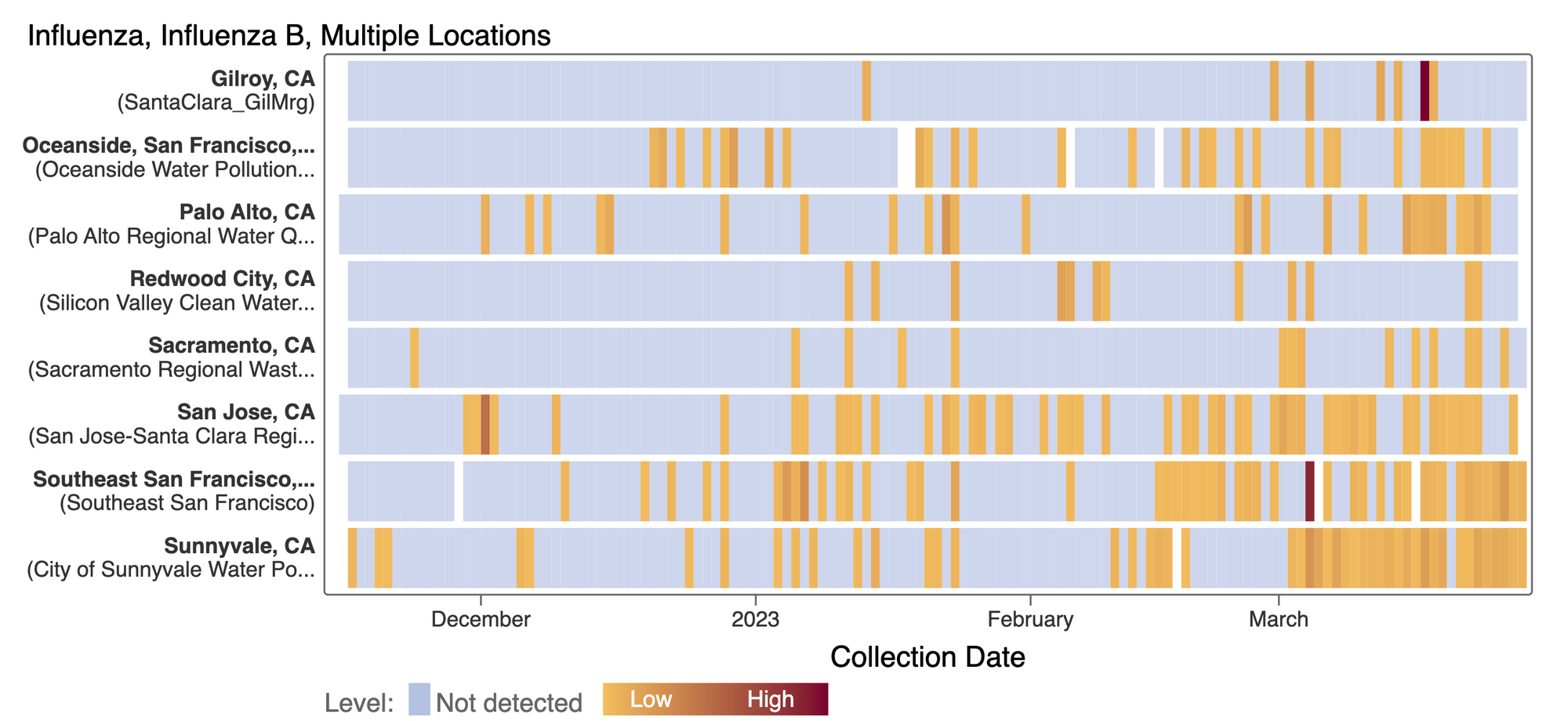
Influenza B (IBV) RNA continues to be sporadically detected at low concentrations at the SCAN plants. The frequency of detection of IBV RNA has definitely increased at some plants including San Jose, Sunnyvale, and Southeast. However, concentrations are still low (less than 5000 copies/g). A heat map is shown below of the IBV RNA concentrations at the SCAN plants. Each plant is represented by a row (y-axis), and dates are shown along the x-axis. Blue indicates a non-detect, and darker orange/red is proportional to concentration with the highest values being darkest. A white means no sample was collected. Here is the link to the heat map chart - the linked chart will update automatically as more data are added to the site.
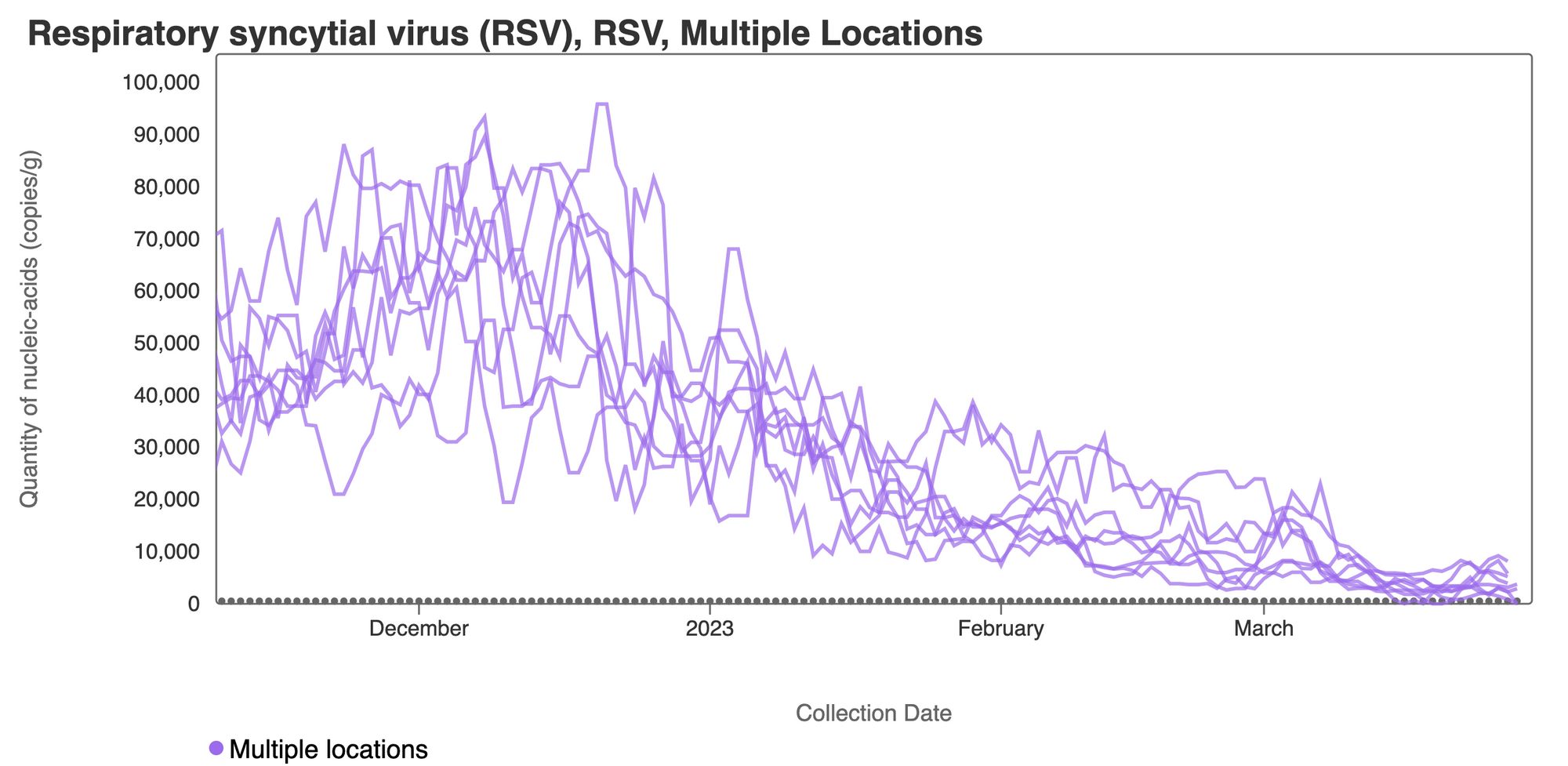
RSV RNA concentrations have decreased from their highest levels but concentrations remain detectable at concentrations between 1,000 - 10,000 copies/gram on all SCAN sites. This plot shows data from all the SCAN sites together since early December 2022 and can be access here. This suggests that there are still individuals shedding RSV RNA into the wastewater in the SCAN sewersheds.
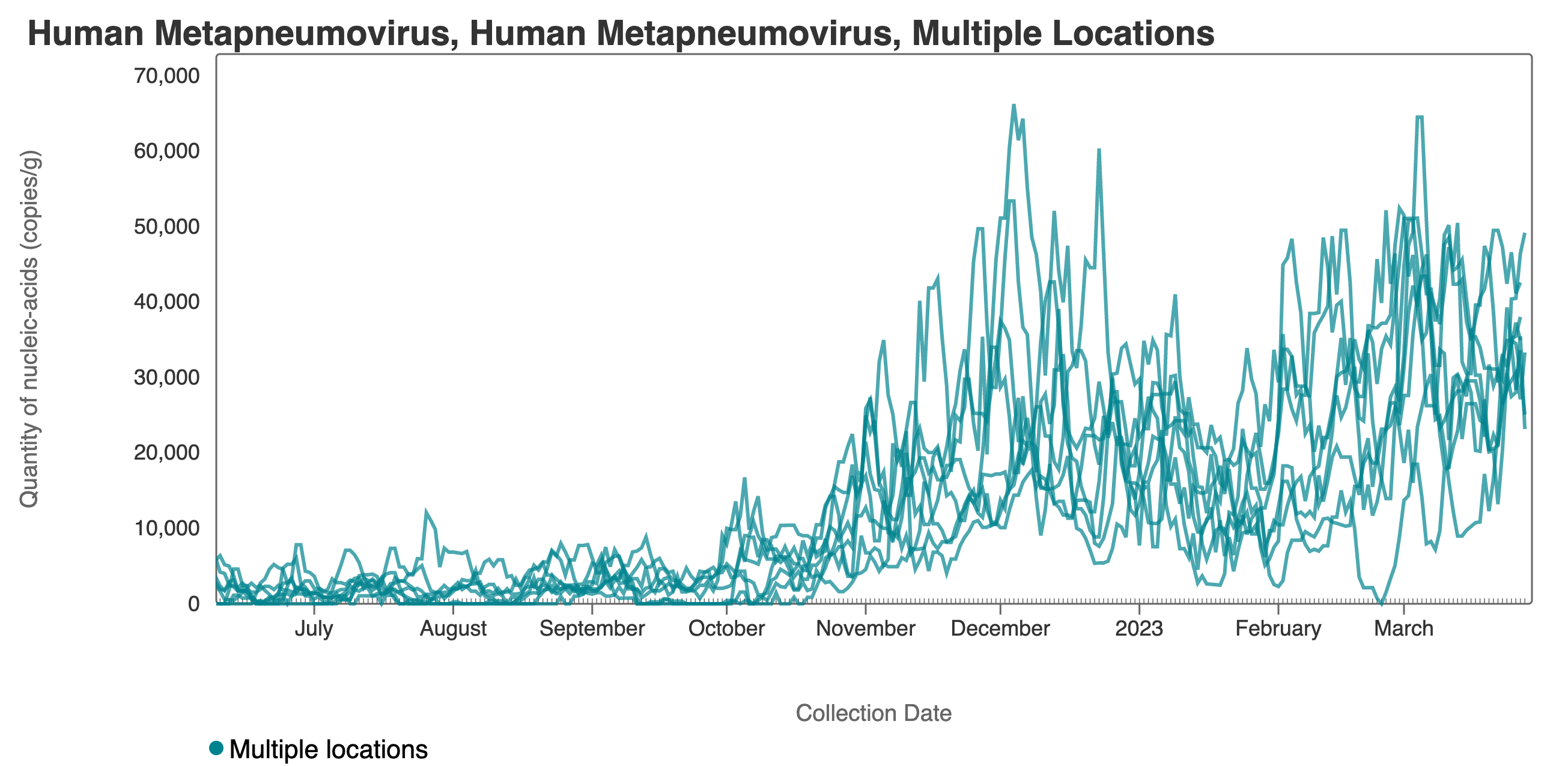
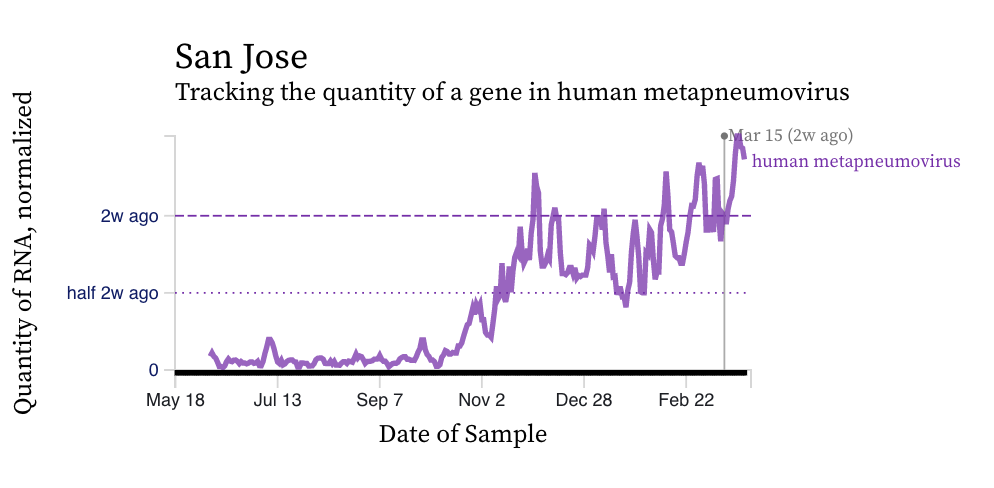
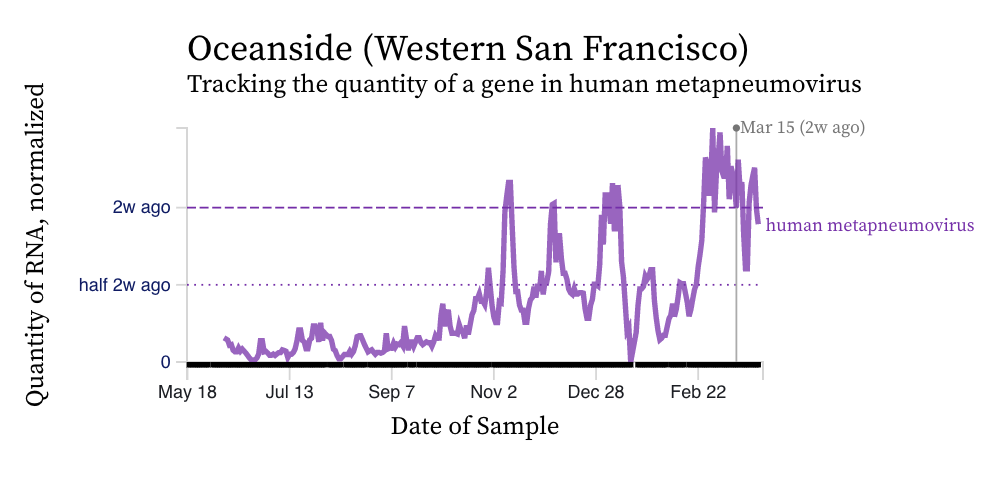
HMPV (human metapneumovirus) RNA concentrations have increased at a number of plants over the last 2 weeks, yet have remained constant at others. Concentrations are between 10,000 and 100,000 copies/g dry weight. Below are charts showing data from all the SCAN plants together (link here to this chart if you want to interact with it), and just San Jose, and just Oceanside. The pattern here is quite distinct from influenza and RSV and suggests this virus is still circulating in the communities.
Norovirus
Assay detects human norovirus GII
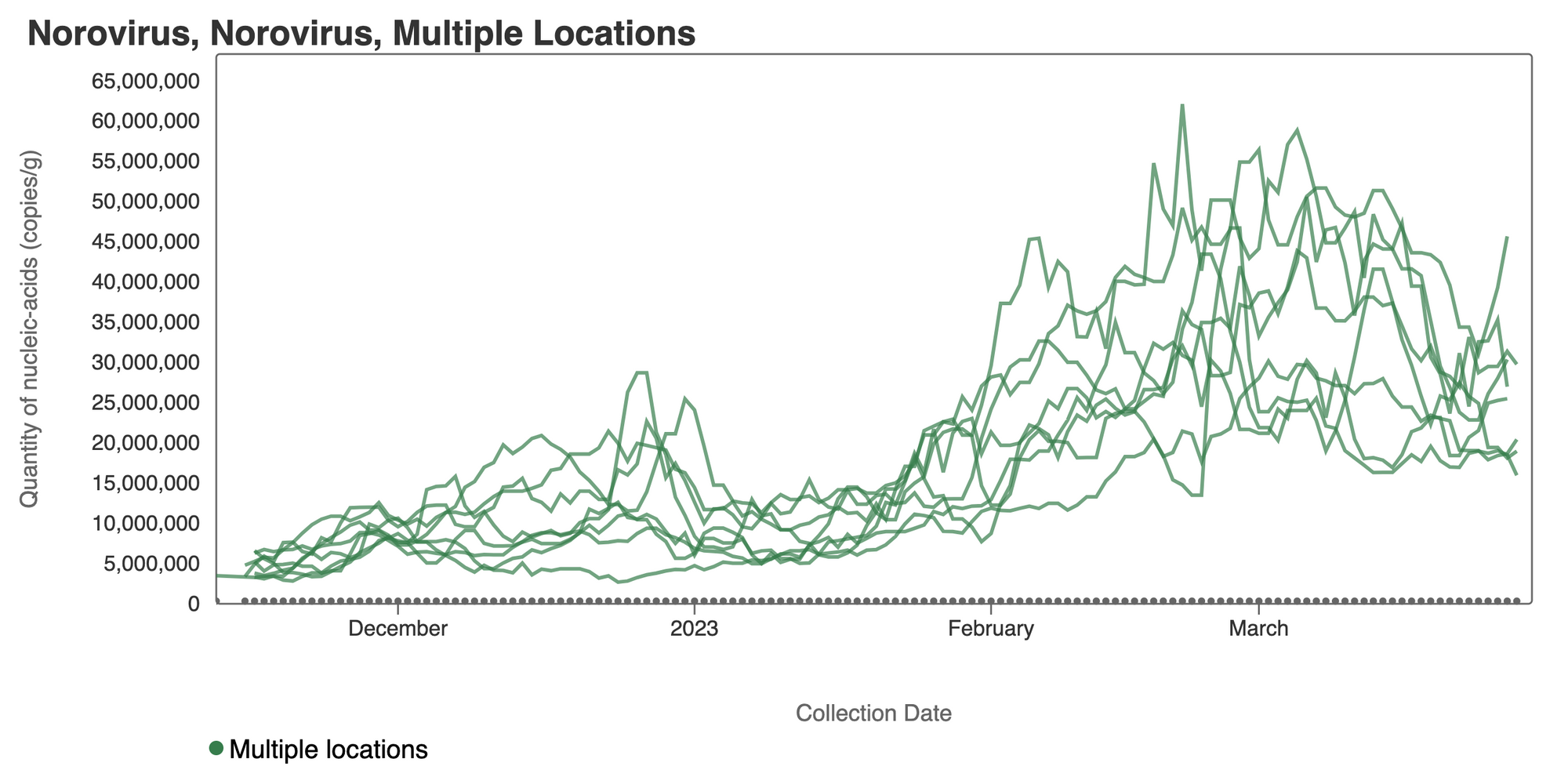
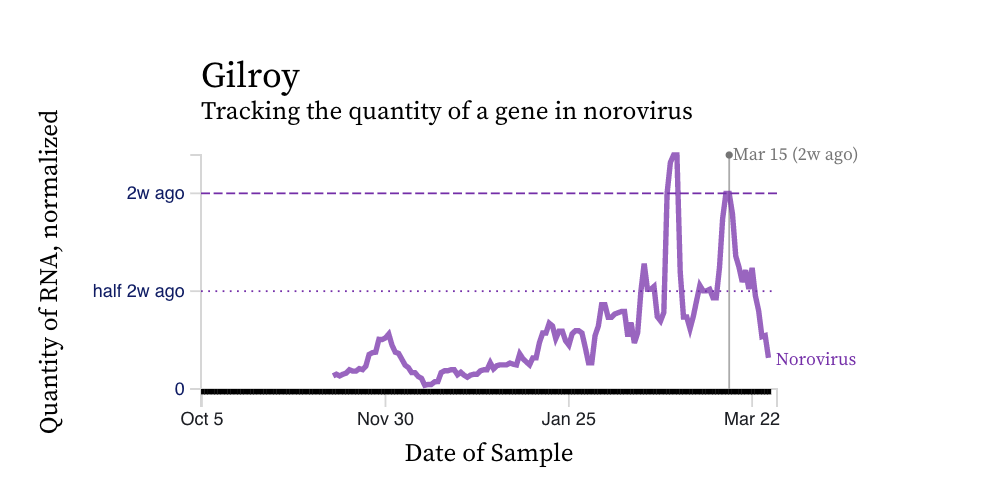
Norovirus GII RNA concentrations have taken a down turn at some plants. A chart of all the SCAN plants together is shown below, and then just Gilroy where a decreasing trend over the last two weeks is clear. You can interact with the chart of all the plants at this link.
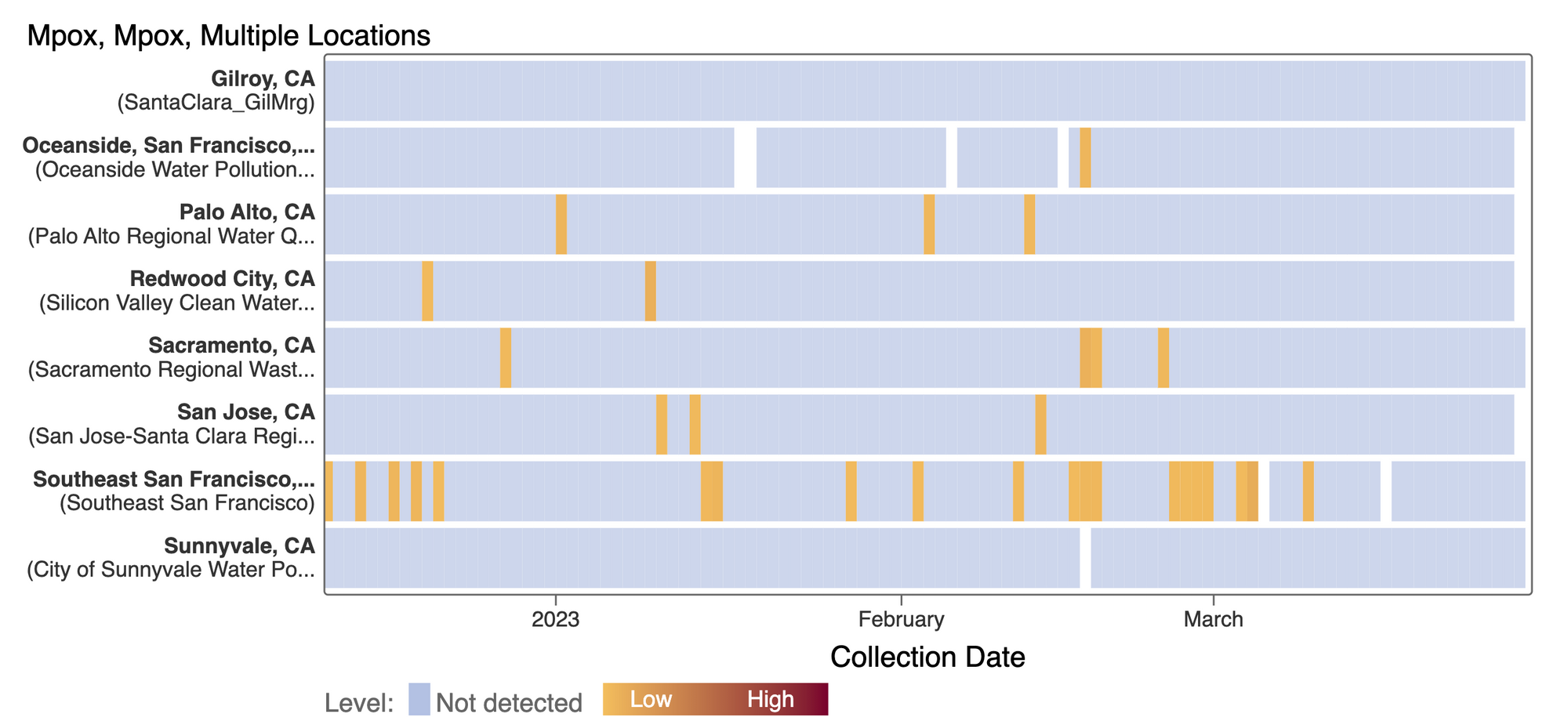
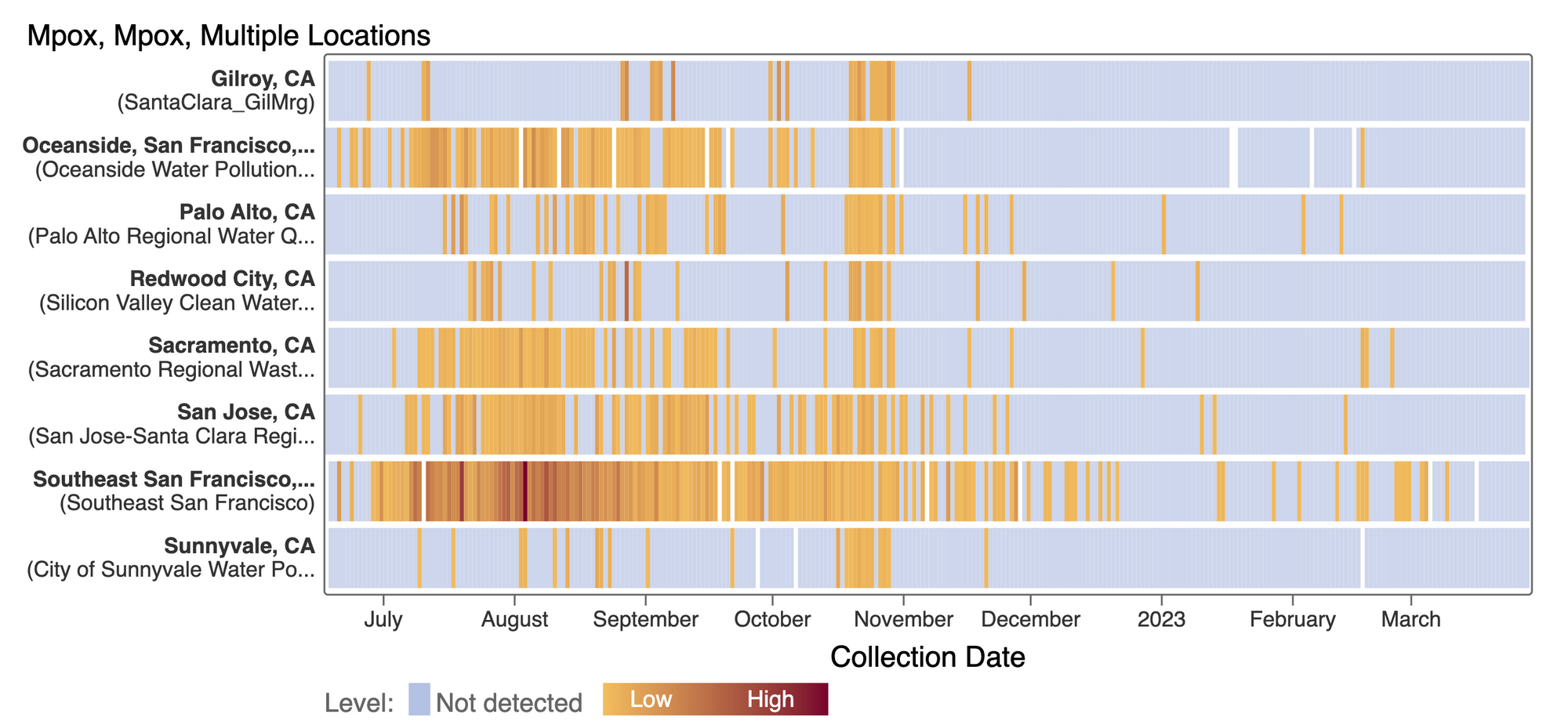
Mpox
We've observed non-detects in the past week for MPXV DNA at the SCAN plants. These heat maps show all the SCAN sites as a row, and each date as a column. The color blue means the sample was non-detect for MPXV DNA and the colors get darker with higher concentrations. The top chart shows just recent data, and the bottom one all the data we have collected, almost a year's worth of data. The charts highlight that we rarely have detections of MPXV DNA at these SCAN plants recently, but we can see how we had much more in the past. You can access this plot here.